Working with missing data
In this section, we will discuss missing (also referred to as NA) values in pandas.
Note The choice of using
NaNinternally to denote missing data was largely for simplicity and performance reasons. Starting from pandas 1.0, some optional data types start experimenting with a nativeNAscalar using a mask-based approach. See here for more.
See the cookbook for some advanced strategies.
Values considered “missing”
As data comes in many shapes and forms, pandas aims to be flexible with regard to handling missing data. While NaN is the default missing value marker for reasons of computational speed and convenience, we need to be able to easily detect this value with data of different types: floating point, integer, boolean, and general object. In many cases, however, the Python None will arise and we wish to also consider that “missing” or “not available” or “NA”.
Note
If you want to considerinfand-infto be “NA” in computations, you can set `pandas.options.mode. use_inf_as_na = True.
In [1]: df = pd.DataFrame(np.random.randn(5, 3),
index=['a', 'c', 'e', 'f', 'h'],
columns=['one', 'two', 'three'])
In [2]: df['four'] = 'bar'
In [3]: df['five'] = df['one'] > 0
In [4]: df
#Out[4]:
one two three four five
a 0.469112 -0.282863 -1.509059 bar True
c -1.135632 1.212112 -0.173215 bar False
e 0.119209 -1.044236 -0.861849 bar True
f -2.104569 -0.494929 1.071804 bar False
h 0.721555 -0.706771 -1.039575 bar True
In [5]: df2 = df.reindex(['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h'])
In [6]: df2
#Out[6]:
one two three four five
a 0.469112 -0.282863 -1.509059 bar True
b NaN NaN NaN NaN NaN
c -1.135632 1.212112 -0.173215 bar False
d NaN NaN NaN NaN NaN
e 0.119209 -1.044236 -0.861849 bar True
f -2.104569 -0.494929 1.071804 bar False
g NaN NaN NaN NaN NaN
h 0.721555 -0.706771 -1.039575 bar True
To make detecting missing values easier (and across different array dtypes), pandas provides the isna() and notna() functions, which are also methods on Series and DataFrame objects:
In [7]: df2['one']
#Out[7]:
a 0.469112
b NaN
c -1.135632
d NaN
e 0.119209
f -2.104569
g NaN
h 0.721555
Name: one, dtype: float64
In [8]: pd.isna(df2['one'])
#Out[8]:
a False
b True
c False
d True
e False
f False
g True
h False
Name: one, dtype: bool
In [9]: df2['four'].notna()
#Out[9]:
a True
b False
c True
d False
e True
f True
g False
h True
Name: four, dtype: bool
In [10]: df2.isna()
#Out[11]:
one two three four five
a False False False False False
b True True True True True
c False False False False False
d True True True True True
e False False False False False
f False False False False False
g True True True True True
h False False False False False
Warning One has to be mindful that in Python (and NumPy), the
nan'sdon’t compare equal, butNone'sdo. Note that pandas/NumPy uses the fact thatnp.nan != np.nan, and treatsNonelikenp.nan.
In [11]: None == None # noqa: E711
#Out[11]: True
In [12]: np.nan == np.nan
#Out[12]: False
So as compared to above, a scalar equality comparison versus a None/np.nan doesn’t provide useful information.
In [13]: df2['one'] == np.nan
a False
b False
c False
d False
e False
f False
g False
h False
Name: one, dtype: bool
Integer dtypes and missing data
Because NaN is a float, a column of integers with even one missing values is cast to floating-point dtype (see Support for integer NA for more). Pandas provides a nullable integer array, which can be used by explicitly requesting the dtype:
In [14]: pd.Series([1, 2, np.nan, 4], dtype=pd.Int64Dtype())
#Out[14]:
0 1
1 2
2 <NA>
3 4
dtype: Int64
Alternatively, the string alias dtype='Int64' (note the capital "I") can be used.
See Nullable integer data type for more.
Datetimes
For datetime64[ns] types, NaT represents missing values. This is a pseudo-native sentinel value that can be represented by NumPy in a singular dtype (datetime64[ns]). pandas objects provide compatibility between NaT and NaN.
In [15]: df2 = df.copy()
In [16]: df2['timestamp'] = pd.Timestamp('20120101')
In [17]: df2
#Out[17]:
one two three four five timestamp
a 0.469112 -0.282863 -1.509059 bar True 2012-01-01
c -1.135632 1.212112 -0.173215 bar False 2012-01-01
e 0.119209 -1.044236 -0.861849 bar True 2012-01-01
f -2.104569 -0.494929 1.071804 bar False 2012-01-01
h 0.721555 -0.706771 -1.039575 bar True 2012-01-01
In [18]: df2.loc[['a', 'c', 'h'], ['one', 'timestamp']] = np.nan
In [19]: df2
#Out[19]:
one two three four five timestamp
a NaN -0.282863 -1.509059 bar True NaT
c NaN 1.212112 -0.173215 bar False NaT
e 0.119209 -1.044236 -0.861849 bar True 2012-01-01
f -2.104569 -0.494929 1.071804 bar False 2012-01-01
h NaN -0.706771 -1.039575 bar True NaT
In [20]: df2.dtypes.value_counts()
#Out[20]:
float64 3
datetime64[ns] 1
bool 1
object 1
dtype: int64
Inserting missing data
You can insert missing values by simply assigning to containers. The actual missing value used will be chosen based on the dtype.
For example, numeric containers will always use NaN regardless of the missing value type chosen:
In [21]: s = pd.Series([1, 2, 3])
In [22]: s.loc[0] = None
In [23]: s
#Out[23]:
0 NaN
1 2.0
2 3.0
dtype: float64
Likewise, datetime containers will always use NaT.
For object containers, pandas will use the value given:
In [24]: s = pd.Series(["a", "b", "c"])
In [25]: s.loc[0] = None
In [26]: s.loc[1] = np.nan
In [27]: s
#Out[27]:
0 None
1 NaN
2 c
dtype: object
Calculations with missing data
Missing values propagate naturally through arithmetic operations between pandas objects.
In [28]: a
#Out[28]:
one two
a NaN -0.282863
c NaN 1.212112
e 0.119209 -1.044236
f -2.104569 -0.494929
h -2.104569 -0.706771
In [29]: b
#Out[29]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e 0.119209 -1.044236 -0.861849
f -2.104569 -0.494929 1.071804
h NaN -0.706771 -1.039575
In [30]: a + b
#Out[30]:
one three two
a NaN NaN -0.565727
c NaN NaN 2.424224
e 0.238417 NaN -2.088472
f -4.209138 NaN -0.989859
h NaN NaN -1.413542
The descriptive statistics and computational methods discussed in the data structure overview (and listed here and here) are all written to account for missing data. For example:
When summing data, NA (missing) values will be treated as zero.
If the data are all NA, the result will be 0.
Cumulative methods like
cumsum()andcumprod()ignore NA values by default, but preserve them in the resulting arrays. To override this behaviour and include NA values, useskipna=False.In [31]: df #Out[31]: one two three a NaN -0.282863 -1.509059 c NaN 1.212112 -0.173215 e 0.119209 -1.044236 -0.861849 f -2.104569 -0.494929 1.071804 h NaN -0.706771 -1.039575In [32]: df['one'].sum() #Out[32]: -1.9853605075978744In [33]: df.mean(1) #Out[33]: a -0.895961 c 0.519449 e -0.595625 f -0.509232 h -0.873173 dtype: float64In [34]: df.cumsum() #Out[34]: one two three a NaN -0.282863 -1.509059 c NaN 0.929249 -1.682273 e 0.119209 -0.114987 -2.544122 f -1.985361 -0.609917 -1.472318 h NaN -1.316688 -2.511893In [35]: df.cumsum(skipna=False) #Out[35]: one two three a NaN -0.282863 -1.509059 c NaN 0.929249 -1.682273 e NaN -0.114987 -2.544122 f NaN -0.609917 -1.472318 h NaN -1.316688 -2.511893
Sum/prod of empties/nans
Warning This behavior is now standard as of v0.22.0 and is consistent with the default in
numpy; previously sum/prod of all-NA or empty Series/DataFrames would return NaN. See v0.22.0 whatsnew for more.
The sum of an empty or all-NA Series or column of a DataFrame is 0.
In [36]: pd.Series([np.nan]).sum()
#Out[36]: 0.0
In [37]: pd.Series([], dtype="float64").sum()
#Out[37]: 0.0
The product of an empty or all-NA Series or column of a DataFrame is 1.
In [38]: pd.Series([np.nan]).prod()
#Out[38]: 1.0
In [39]: pd.Series([], dtype="float64").prod()
#Out[39]: 1.0
NA values in GroupBy
NA groups in GroupBy are automatically excluded. This behavior is consistent with R, for example:
In [40]: df
#Out[40]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e 0.119209 -1.044236 -0.861849
f -2.104569 -0.494929 1.071804
h NaN -0.706771 -1.039575
In [41]: df.groupby('one').mean()
#Out[41]:
two three
one
-2.104569 -0.494929 1.071804
0.119209 -1.044236 -0.861849
See the groupby section here for more information.
Cleaning / filling missing data[](https://pandas.pydata.org/pandas-docs/stable/user_guide/missing_data.html#cleaning-filling-missing-data “Permalink to this headline”)
pandas objects are equipped with various data manipulation methods for dealing with missing data.
Filling missing values: fillna
fillna() can “fill in” NA values with non-NA data in a couple of ways, which we illustrate:
Replace NA with a scalar value
In [42]: df2
#Out[42]:
one two three four five timestamp
a NaN -0.282863 -1.509059 bar True NaT
c NaN 1.212112 -0.173215 bar False NaT
e 0.119209 -1.044236 -0.861849 bar True 2012-01-01
f -2.104569 -0.494929 1.071804 bar False 2012-01-01
h NaN -0.706771 -1.039575 bar True NaT
In [43]: df2.fillna(0)
#Out[43]:
one two three four five timestamp
a 0.000000 -0.282863 -1.509059 bar True 0
c 0.000000 1.212112 -0.173215 bar False 0
e 0.119209 -1.044236 -0.861849 bar True 2012-01-01 00:00:00
f -2.104569 -0.494929 1.071804 bar False 2012-01-01 00:00:00
h 0.000000 -0.706771 -1.039575 bar True 0
In [44]: df2['one'].fillna('missing')
#Out[44]:
a missing
c missing
e 0.119209
f -2.10457
h missing
Name: one, dtype: object
Fill gaps forward or backward
Using the same filling arguments as reindexing, we can propagate non-NA values forward or backward:
In [45]: df
#Out[45]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e 0.119209 -1.044236 -0.861849
f -2.104569 -0.494929 1.071804
h NaN -0.706771 -1.039575
In [46]: df.fillna(method='pad')
#Out[46]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e 0.119209 -1.044236 -0.861849
f -2.104569 -0.494929 1.071804
h -2.104569 -0.706771 -1.039575
Limit the amount of filling
If we only want consecutive gaps filled up to a certain number of data points, we can use the limit keyword:
In [47]: df
#Out[47]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e NaN NaN NaN
f NaN NaN NaN
h NaN -0.706771 -1.039575
In [48]: df.fillna(method='pad', limit=1)
#Out[48]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e NaN 1.212112 -0.173215
f NaN NaN NaN
h NaN -0.706771 -1.039575
To remind you, these are the available filling methods:
| Method | Action |
|---|---|
| pad / ffill | Fill values forward |
| bfill / backfill | Fill values backward |
With time series data, using pad/ffill is extremely common so that the “last known value” is available at every time point.
ffill() is equivalent to fillna(method='ffill') and bfill() is equivalent to fillna(method='bfill')
Filling with a PandasObject
You can also fillna using a dict or Series that is alignable. The labels of the dict or index of the Series must match the columns of the frame you wish to fill. The use case of this is to fill a DataFrame with the mean of that column.
In [49]: dff = pd.DataFrame(np.random.randn(10, 3), columns=list('ABC'))
In [50]: dff.iloc[3:5, 0] = np.nan
In [51]: dff.iloc[4:6, 1] = np.nan
In [52]: dff.iloc[5:8, 2] = np.nan
In [53]: dff
#Out[53]:
A B C
0 0.271860 -0.424972 0.567020
1 0.276232 -1.087401 -0.673690
2 0.113648 -1.478427 0.524988
3 NaN 0.577046 -1.715002
4 NaN NaN -1.157892
5 -1.344312 NaN NaN
6 -0.109050 1.643563 NaN
7 0.357021 -0.674600 NaN
8 -0.968914 -1.294524 0.413738
9 0.276662 -0.472035 -0.013960
In [54]: dff.fillna(dff.mean())
#Out[54]:
A B C
0 0.271860 -0.424972 0.567020
1 0.276232 -1.087401 -0.673690
2 0.113648 -1.478427 0.524988
3 -0.140857 0.577046 -1.715002
4 -0.140857 -0.401419 -1.157892
5 -1.344312 -0.401419 -0.293543
6 -0.109050 1.643563 -0.293543
7 0.357021 -0.674600 -0.293543
8 -0.968914 -1.294524 0.413738
9 0.276662 -0.472035 -0.013960
In [55]: dff.fillna(dff.mean()['B':'C'])
#Out[55]:
A B C
0 0.271860 -0.424972 0.567020
1 0.276232 -1.087401 -0.673690
2 0.113648 -1.478427 0.524988
3 NaN 0.577046 -1.715002
4 NaN -0.401419 -1.157892
5 -1.344312 -0.401419 -0.293543
6 -0.109050 1.643563 -0.293543
7 0.357021 -0.674600 -0.293543
8 -0.968914 -1.294524 0.413738
9 0.276662 -0.472035 -0.013960
Same result as above, but is aligning the ‘fill’ value which is a Series in this case.
In [56]: dff.where(pd.notna(dff), dff.mean(), axis='columns')
#Out[56]:
A B C
0 0.271860 -0.424972 0.567020
1 0.276232 -1.087401 -0.673690
2 0.113648 -1.478427 0.524988
3 -0.140857 0.577046 -1.715002
4 -0.140857 -0.401419 -1.157892
5 -1.344312 -0.401419 -0.293543
6 -0.109050 1.643563 -0.293543
7 0.357021 -0.674600 -0.293543
8 -0.968914 -1.294524 0.413738
9 0.276662 -0.472035 -0.013960
Dropping axis labels with missing data: dropna[](https://pandas.pydata.org/pandas-docs/stable/user_guide/missing_data.html#dropping-axis-labels-with-missing-data-dropna “Permalink to this headline”)
You may wish to simply exclude labels from a data set which refer to missing data. To do this, use dropna():
In [57]: df
#Out[57]:
one two three
a NaN -0.282863 -1.509059
c NaN 1.212112 -0.173215
e NaN 0.000000 0.000000
f NaN 0.000000 0.000000
h NaN -0.706771 -1.039575
In [58]: df.dropna(axis=0)
#Out[58]:
Empty DataFrame
Columns: [one, two, three]
Index: []
In [59]: df.dropna(axis=1)
#Out[59]:
two three
a -0.282863 -1.509059
c 1.212112 -0.173215
e 0.000000 0.000000
f 0.000000 0.000000
h -0.706771 -1.039575
In [60]: df['one'].dropna()
#Out[60]: Series([], Name: one, dtype: float64)
An equivalent dropna() is available for Series. DataFrame.dropna has considerably more options than Series.dropna, which can be examined in the API.
Interpolation
New in version 0.23.0: The limit_area keyword argument was added.
Both Series and DataFrame objects have interpolate() that, by default, performs linear interpolation at missing data points.
In [61]: ts
#Out[61]:
2000-01-31 0.469112
2000-02-29 NaN
2000-03-31 NaN
2000-04-28 NaN
2000-05-31 NaN
...
2007-12-31 -6.950267
2008-01-31 -7.904475
2008-02-29 -6.441779
2008-03-31 -8.184940
2008-04-30 -9.011531
Freq: BM, Length: 100, dtype: float64
In [62]: ts.count()
#Out[62]: 66
In [63]: ts.plot()
#Out[63]:
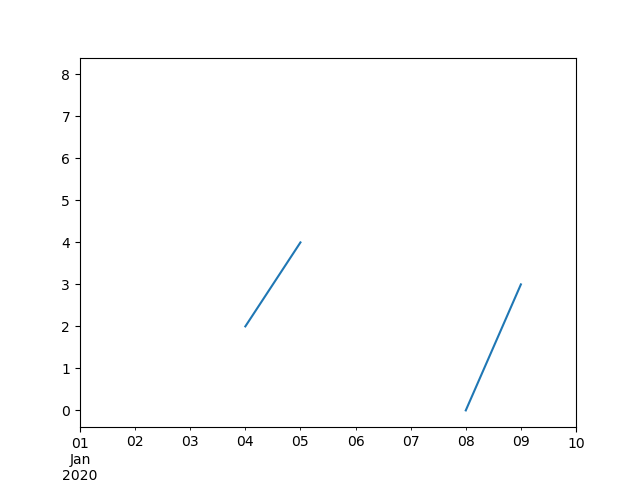
In [64]: ts.interpolate()
#Out[64]:
2000-01-31 0.469112
2000-02-29 0.434469
2000-03-31 0.399826
2000-04-28 0.365184
2000-05-31 0.330541
...
2007-12-31 -6.950267
2008-01-31 -7.904475
2008-02-29 -6.441779
2008-03-31 -8.184940
2008-04-30 -9.011531
Freq: BM, Length: 100, dtype: float64
In [65]: ts.interpolate().count()
#Out[65]: 100
In [66]: ts.interpolate().plot()
#Out[66]:

Index aware interpolation is available via the method keyword:
In [67]: ts2
#Out[67]:
2000-01-31 0.469112
2000-02-29 NaN
2002-07-31 -5.785037
2005-01-31 NaN
2008-04-30 -9.011531
dtype: float64
In [68]: ts2.interpolate()
#Out[68]:
2000-01-31 0.469112
2000-02-29 -2.657962
2002-07-31 -5.785037
2005-01-31 -7.398284
2008-04-30 -9.011531
dtype: float64
In [69]: ts2.interpolate(method='time')
#Out[69]:
2000-01-31 0.469112
2000-02-29 0.270241
2002-07-31 -5.785037
2005-01-31 -7.190866
2008-04-30 -9.011531
dtype: float64
For a floating-point index, use method='values':
In [70]: ser
#Out[70]:
0.0 0.0
1.0 NaN
10.0 10.0
dtype: float64
In [71]: ser.interpolate()
#Out[71]:
0.0 0.0
1.0 5.0
10.0 10.0
dtype: float64
In [72]: ser.interpolate(method='values')
#Out[72]:
0.0 0.0
1.0 1.0
10.0 10.0
dtype: float64
You can also interpolate with a DataFrame:
In [73]: df = pd.DataFrame({'A': [1, 2.1, np.nan, 4.7, 5.6, 6.8],
....: 'B': [.25, np.nan, np.nan, 4, 12.2, 14.4]})
....:
In [74]: df
#Out[74]:
A B
0 1.0 0.25
1 2.1 NaN
2 NaN NaN
3 4.7 4.00
4 5.6 12.20
5 6.8 14.40
In [75]: df.interpolate()
#Out[75]:
A B
0 1.0 0.25
1 2.1 1.50
2 3.4 2.75
3 4.7 4.00
4 5.6 12.20
5 6.8 14.40
The method argument gives access to fancier interpolation methods. If you have scipy installed, you can pass the name of a 1-d interpolation r#Outine to method. You’ll want to consult the full scipy interpolation documentation and reference guide for details. The appropriate interpolation method will depend on the type of data you are working with.
If you are dealing with a time series that is growing at an increasing rate,
method='quadratic'may be appropriate.If you have values approximating a cumulative distribution function, then
method='pchip'should work well.To fill missing values with goal of smooth plotting, consider
method='akima'.
Warning
These methods require scipy.
In [76]: df.interpolate(method='barycentric')
#Out[76]:
A B
0 1.00 0.250
1 2.10 -7.660
2 3.53 -4.515
3 4.70 4.000
4 5.60 12.200
5 6.80 14.400
In [77]: df.interpolate(method='pchip')
#Out[77]:
A B
0 1.00000 0.250000
1 2.10000 0.672808
2 3.43454 1.928950
3 4.70000 4.000000
4 5.60000 12.200000
5 6.80000 14.400000
In [78]: df.interpolate(method='akima')
#Out[78]:
A B
0 1.000000 0.250000
1 2.100000 -0.873316
2 3.406667 0.320034
3 4.700000 4.000000
4 5.600000 12.200000
5 6.800000 14.400000
When interpolating via a polynomial or spline approximation, you must also specify the degree or order of the approximation:
In [79]: df.interpolate(method='spline', order=2)
#Out[79]:
A B
0 1.000000 0.250000
1 2.100000 -0.428598
2 3.404545 1.206900
3 4.700000 4.000000
4 5.600000 12.200000
5 6.800000 14.400000
In [80]: df.interpolate(method='polynomial', order=2)
#Out[80]:
A B
0 1.000000 0.250000
1 2.100000 -2.703846
2 3.451351 -1.453846
3 4.700000 4.000000
4 5.600000 12.200000
5 6.800000 14.400000
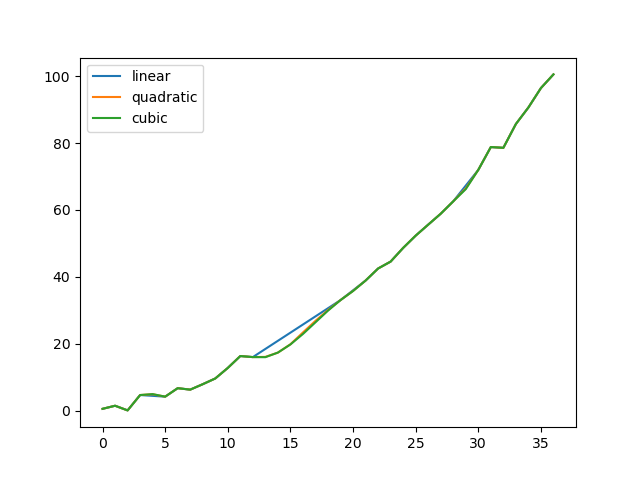
Compare several methods:
In [81]: np.random.seed(2)
In [82]: ser = pd.Series(np.arange(1, 10.1, .25) ** 2 + np.random.randn(37))
In [83]: missing = np.array([4, 13, 14, 15, 16, 17, 18, 20, 29])
In [84]: ser[missing] = np.nan
In [85]: methods = ['linear', 'quadratic', 'cubic']
In [86]: df = pd.DataFrame({m: ser.interpolate(method=m) for m in methods})
In [87]: df.plot()
#Out[87]:
Another use case is interpolation at new values. Suppose you have 100 observations from some distribution. And let’s suppose that you’re particularly interested in what’s happening around the middle. You can mix pandas’ reindex and interpolate methods to interpolate at the new values.
In [88]: ser = pd.Series(np.sort(np.random.uniform(size=100)))
# interpolate at new_index
In [89]: new_index = ser.index | pd.Index([49.25, 49.5, 49.75, 50.25, 50.5, 50.75])
In [90]: interp_s = ser.reindex(new_index).interpolate(method='pchip')
In [91]: interp_s[49:51]
#Out[91]:
49.00 0.471410
49.25 0.476841
49.50 0.481780
49.75 0.485998
50.00 0.489266
50.25 0.491814
50.50 0.493995
50.75 0.495763
51.00 0.497074
dtype: float64
Interpolation limits[](https://pandas.pydata.org/pandas-docs/stable/user_guide/missing_data.html#interpolation-limits “Permalink to this headline”)
Like other pandas fill methods, interpolate() accepts a limit keyword argument. Use this argument to limit the number of consecutive NaN values filled since the last valid observation:
In [92]: ser = pd.Series([np.nan, np.nan, 5, np.nan, np.nan,
....: np.nan, 13, np.nan, np.nan])
....:
In [93]: ser
#Out[93]:
0 NaN
1 NaN
2 5.0
3 NaN
4 NaN
5 NaN
6 13.0
7 NaN
8 NaN
dtype: float64
# fill all consecutive values in a forward direction
In [94]: ser.interpolate()
#Out[94]:
0 NaN
1 NaN
2 5.0
3 7.0
4 9.0
5 11.0
6 13.0
7 13.0
8 13.0
dtype: float64
# fill one consecutive value in a forward direction
In [95]: ser.interpolate(limit=1)
#Out[95]:
0 NaN
1 NaN
2 5.0
3 7.0
4 NaN
5 NaN
6 13.0
7 13.0
8 NaN
dtype: float64
By default, NaN values are filled in a forward direction. Use limit_direction parameter to fill backward or from both directions.
# fill one consecutive value backwards
In [96]: ser.interpolate(limit=1, limit_direction='backward')
#Out[96]:
0 NaN
1 5.0
2 5.0
3 NaN
4 NaN
5 11.0
6 13.0
7 NaN
8 NaN
dtype: float64
# fill one consecutive value in both directions
In [97]: ser.interpolate(limit=1, limit_direction='both')
#Out[97]:
0 NaN
1 5.0
2 5.0
3 7.0
4 NaN
5 11.0
6 13.0
7 13.0
8 NaN
dtype: float64
# fill all consecutive values in both directions
In [98]: ser.interpolate(limit_direction='both')
#Out[98]:
0 5.0
1 5.0
2 5.0
3 7.0
4 9.0
5 11.0
6 13.0
7 13.0
8 13.0
dtype: float64
By default, NaN values are filled whether they are inside (surrounded by) existing valid values, or #Outside existing valid values. Introduced in v0.23 the limit_area parameter restricts filling to either inside or #Outside values.
# fill one consecutive inside value in both directions
In [99]: ser.interpolate(limit_direction='both', limit_area='inside', limit=1)
#Out[99]:
0 NaN
1 NaN
2 5.0
3 7.0
4 NaN
5 11.0
6 13.0
7 NaN
8 NaN
dtype: float64
# fill all consecutive #Outside values backward
In [100]: ser.interpolate(limit_direction='backward', limit_area='#Outside')
#Out[100]:
0 5.0
1 5.0
2 5.0
3 NaN
4 NaN
5 NaN
6 13.0
7 NaN
8 NaN
dtype: float64
# fill all consecutive #Outside values in both directions
In [101]: ser.interpolate(limit_direction='both', limit_area='#Outside')
#Out[101]:
0 5.0
1 5.0
2 5.0
3 NaN
4 NaN
5 NaN
6 13.0
7 13.0
8 13.0
dtype: float64
Replacing generic values
Often times we want to replace arbitrary values with other values.
replace() in Series and replace() in DataFrame provides an efficient yet flexible way to perform such replacements.
For a Series, you can replace a single value or a list of values by another value:
In [102]: ser = pd.Series([0., 1., 2., 3., 4.])
In [103]: ser.replace(0, 5)
#Out[103]:
0 5.0
1 1.0
2 2.0
3 3.0
4 4.0
dtype: float64
You can replace a list of values by a list of other values:
In [104]: ser.replace([0, 1, 2, 3, 4], [4, 3, 2, 1, 0])
#Out[104]:
0 4.0
1 3.0
2 2.0
3 1.0
4 0.0
dtype: float64
You can also specify a mapping dict:
In [105]: ser.replace({0: 10, 1: 100})
#Out[105]:
0 10.0
1 100.0
2 2.0
3 3.0
4 4.0
dtype: float64
For a DataFrame, you can specify individual values by column:
In [106]: df = pd.DataFrame({'a': [0, 1, 2, 3, 4], 'b': [5, 6, 7, 8, 9]})
In [107]: df.replace({'a': 0, 'b': 5}, 100)
#Out[107]:
a b
0 100 100
1 1 6
2 2 7
3 3 8
4 4 9
Instead of replacing with specified values, you can treat all given values as missing and interpolate over them:
In [108]: ser.replace([1, 2, 3], method='pad')
#Out[108]:
0 0.0
1 0.0
2 0.0
3 0.0
4 4.0
dtype: float64
String/regular expression replacement
Note
Python strings prefixed with the r character such as r'hello world' are so-called “raw” strings. They have different semantics regarding backslashes than strings with#Out this prefix. Backslashes in raw strings will be interpreted as an escaped backslash, e.g., r'\' == '\\'. You should read ab#Out them if this is unclear.
Replace the ‘.’ with NaN (str -> str):
In [109]: d = {'a': list(range(4)), 'b': list('ab..'), 'c': ['a', 'b', np.nan, 'd']}
In [110]: df = pd.DataFrame(d)
In [111]: df.replace('.', np.nan)
#Out[111]:
a b c
0 0 a a
1 1 b b
2 2 NaN NaN
3 3 NaN d
Now do it with a regular expression that removes surrounding whitespace (regex -> regex):
In [112]: df.replace(r'\s*\.\s*', np.nan, regex=True)
#Out[112]:
a b c
0 0 a a
1 1 b b
2 2 NaN NaN
3 3 NaN d
Replace a few different values (list -> list):
In [113]: df.replace(['a', '.'], ['b', np.nan])
#Out[113]:
a b c
0 0 b b
1 1 b b
2 2 NaN NaN
3 3 NaN d
list of regex -> list of regex:
In [114]: df.replace([r'\.', r'(a)'], ['dot', r'\1stuff'], regex=True)
#Out[114]:
a b c
0 0 astuff astuff
1 1 b b
2 2 dot NaN
3 3 dot d
Only search in column 'b' (dict -> dict):
In [115]: df.replace({'b': '.'}, {'b': np.nan})
#Out[115]:
a b c
0 0 a a
1 1 b b
2 2 NaN NaN
3 3 NaN d
Same as the previous example, but use a regular expression for searching instead (dict of regex -> dict):
In [116]: df.replace({'b': r'\s*\.\s*'}, {'b': np.nan}, regex=True)
#Out[116]:
a b c
0 0 a a
1 1 b b
2 2 NaN NaN
3 3 NaN d
You can pass nested dictionaries of regular expressions that use regex=True:
In [117]: df.replace({'b': {'b': r''}}, regex=True)
#Out[117]:
a b c
0 0 a a
1 1 b
2 2 . NaN
3 3 . d
Alternatively, you can pass the nested dictionary like so:
In [118]: df.replace(regex={'b': {r'\s*\.\s*': np.nan}})
#Out[118]:
a b c
0 0 a a
1 1 b b
2 2 NaN NaN
3 3 NaN d
You can also use the group of a regular expression match when replacing (dict of regex -> dict of regex), this works for lists as well.
In [119]: df.replace({'b': r'\s*(\.)\s*'}, {'b': r'\1ty'}, regex=True)
#Out[119]:
a b c
0 0 a a
1 1 b b
2 2 .ty NaN
3 3 .ty d
You can pass a list of regular expressions, of which those that match will be replaced with a scalar (list of regex -> regex).
In [120]: df.replace([r'\s*\.\s*', r'a|b'], np.nan, regex=True)
#Out[120]:
a b c
0 0 NaN NaN
1 1 NaN NaN
2 2 NaN NaN
3 3 NaN d
All of the regular expression examples can also be passed with the to_replace argument as the regex argument. In this case the value argument must be passed explicitly by name or regex must be a nested dictionary. The previous example, in this case, would then be:
In [121]: df.replace(regex=[r'\s*\.\s*', r'a|b'], value=np.nan)
#Out[121]:
a b c
0 0 NaN NaN
1 1 NaN NaN
2 2 NaN NaN
3 3 NaN d
This can be convenient if you do not want to pass regex=True every time you want to use a regular expression.
Note
Anywhere in the above replace examples that you see a regular expression a compiled regular expression is valid as well.
Numeric replacement
replace() is similar to fillna().
In [122]: df = pd.DataFrame(np.random.randn(10, 2))
In [123]: df[np.random.rand(df.shape[0]) > 0.5] = 1.5
In [124]: df.replace(1.5, np.nan)
#Out[124]:
0 1
0 -0.844214 -1.021415
1 0.432396 -0.323580
2 0.423825 0.799180
3 1.262614 0.751965
4 NaN NaN
5 NaN NaN
6 -0.498174 -1.060799
7 0.591667 -0.183257
8 1.019855 -1.482465
9 NaN NaN
Replacing more than one value is possible by passing a list.
In [125]: df00 = df.iloc[0, 0]
In [126]: df.replace([1.5, df00], [np.nan, 'a'])
#Out[126]:
0 1
0 a -1.02141
1 0.432396 -0.32358
2 0.423825 0.79918
3 1.26261 0.751965
4 NaN NaN
5 NaN NaN
6 -0.498174 -1.0608
7 0.591667 -0.183257
8 1.01985 -1.48247
9 NaN NaN
In [127]: df[1].dtype
#Out[127]: dtype('float64')
You can also operate on the DataFrame in place:
In [128]: df.replace(1.5, np.nan, inplace=True)
Warning
When replacing multiple bool or datetime64 objects, the first argument to replace (to_replace) must match the type of the value being replaced. For example,
s = pd.Series([True, False, True])
s.replace({'a string': 'new value', True: False}) # raises
# TypeError: Cannot compare types 'ndarray(dtype=bool)' and 'str'
will raise a TypeError because one of the dict keys is not of the correct type for replacement.
However, when replacing a single object such as,
In [129]: s = pd.Series([True, False, True])
In [130]: s.replace('a string', 'another string')
#Out[130]:
0 True
1 False
2 True
dtype: bool
the original NDFrame object will be returned untouched. We’re working on unifying this API, but for backwards compatibility reasons we cannot break the latter behavior. See GH6354 for more details.
Missing data casting rules and indexing
While pandas supports storing arrays of integer and boolean type, these types are not capable of storing missing data. Until we can switch to using a native NA type in NumPy, we’ve established some “casting rules”. When a reindexing operation introduces missing data, the Series will be cast according to the rules introduced in the table below. | data type | Cast to | |———–|———–| | integer | float | |boolean|object| |float|no cast| |object|no cast|
For example:
In [131]: s = pd.Series(np.random.randn(5), index=[0, 2, 4, 6, 7])
In [132]: s > 0
#Out[132]:
0 True
2 True
4 True
6 True
7 True
dtype: bool
In [133]: (s > 0).dtype
#Out[133]: dtype('bool')
In [134]: crit = (s > 0).reindex(list(range(8)))
In [135]: crit
#Out[135]:
0 True
1 NaN
2 True
3 NaN
4 True
5 NaN
6 True
7 True
dtype: object
In [136]: crit.dtype
#Out[136]: dtype('O')
Ordinarily NumPy will complain if you try to use an object array (even if it contains boolean values) instead of a boolean array to get or set values from an ndarray (e.g. selecting values based on some criteria). If a boolean vector contains NAs, an exception will be generated:
In [137]: reindexed = s.reindex(list(range(8))).fillna(0)
In [138]: reindexed[crit]
---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
<ipython-input-138-0dac417a4890> in <module>
----> 1 reindexed[crit]
/pandas/pandas/core/series.py in __getitem__(self, key)
905 key = list(key)
906
--> 907 if com.is_bool_indexer(key):
908 key = check_bool_indexer(self.index, key)
909
/pandas/pandas/core/common.py in is_bool_indexer(key)
134 if not lib.is_bool_array(key):
135 if isna(key).any():
--> 136 raise ValueError(na_msg)
137 return False
138 return True
ValueError: cannot mask with array containing NA / NaN values
However, these can be filled in using fillna() and it will work fine:
In [139]: reindexed[crit.fillna(False)]
#Out[139]:
0 0.126504
2 0.696198
4 0.697416
6 0.601516
7 0.003659
dtype: float64
In [140]: reindexed[crit.fillna(True)]
#Out[140]:
0 0.126504
1 0.000000
2 0.696198
3 0.000000
4 0.697416
5 0.000000
6 0.601516
7 0.003659
dtype: float64
Pandas provides a nullable integer dtype, but you must explicitly request it when creating the series or column. Notice that we use a capital “I” in the dtype="Int64".
In [141]: s = pd.Series([0, 1, np.nan, 3, 4], dtype="Int64")
In [142]: s
#Out[142]:
0 0
1 1
2 <NA>
3 3
4 4
dtype: Int64
See Nullable integer data type for more.
Experimental NA scalar to denote missing values
Warning
Experimental: the behaviour of pd.NA can still change with#Out warning.
New in version 1.0.0.
Starting from pandas 1.0, an experimental pd.NA value (singleton) is available to represent scalar missing values. At this moment, it is used in the nullable integer, boolean and dedicated string data types as the missing value indicator.
The goal of pd.NA is provide a “missing” indicator that can be used consistently across data types (instead of np.nan, None or pd.NaT depending on the data type).
For example, when having missing values in a Series with the nullable integer dtype, it will use pd.NA:
In [143]: s = pd.Series([1, 2, None], dtype="Int64")
In [144]: s
#Out[144]:
0 1
1 2
2 <NA>
dtype: Int64
In [145]: s[2]
#Out[145]: <NA>
In [146]: s[2] is pd.NA
#Out[146]: True
Currently, pandas does not yet use those data types by default (when creating a DataFrame or Series, or when reading in data), so you need to specify the dtype explicitly. An easy way to convert to those dtypes is explained here.
Propagation in arithmetic and comparison operations
In general, missing values propagate in operations involving pd.NA. When one of the operands is unknown, the #Outcome of the operation is also unknown.
For example, pd.NA propagates in arithmetic operations, similarly to np.nan:
In [147]: pd.NA + 1
#Out[147]: <NA>
In [148]: "a" * pd.NA
#Out[148]: <NA>
There are a few special cases when the result is known, even when one of the operands is NA.
In [149]: pd.NA ** 0
#Out[149]: 1
In [150]: 1 ** pd.NA
#Out[150]: 1
In equality and comparison operations, pd.NA also propagates. This deviates from the behaviour of np.nan, where comparisons with np.nan always return False.
In [151]: pd.NA == 1
#Out[151]: <NA>
In [152]: pd.NA == pd.NA
#Out[152]: <NA>
In [153]: pd.NA < 2.5
#Out[153]: <NA>
To check if a value is equal to pd.NA, the isna() function can be used:
In [154]: pd.isna(pd.NA)
#Out[154]: True
An exception on this basic propagation rule are reductions (such as the mean or the minimum), where pandas defaults to skipping missing values. See above for more.
Logical operations
For logical operations, pd.NA follows the rules of the three-valued logic (or Kleene logic, similarly to R, SQL and Julia). This logic means to only propagate missing values when it is logically required.
For example, for the logical “or” operation (|), if one of the operands is True, we already know the result will be True, regardless of the other value (so regardless the missing value would be True or False). In this case, pd.NA does not propagate:
In [155]: True | False
#Out[155]: True
In [156]: True | pd.NA
#Out[156]: True
In [157]: pd.NA | True
#Out[157]: True
On the other hand, if one of the operands is False, the result depends on the value of the other operand. Therefore, in this case pd.NA propagates:
In [158]: False | True
#Out[158]: True
In [159]: False | False
#Out[159]: False
In [160]: False | pd.NA
#Out[160]: <NA>
The behaviour of the logical “and” operation (&) can be derived using similar logic (where now pd.NA will not propagate if one of the operands is already False):
In [161]: False & True
#Out[161]: False
In [162]: False & False
#Out[162]: False
In [163]: False & pd.NA
#Out[163]: False
In [164]: True & True
#Out[164]: True
In [165]: True & False
#Out[165]: False
In [166]: True & pd.NA
#Out[166]: <NA>
NA in a boolean context
Since the actual value of an NA is unknown, it is ambiguous to convert NA to a boolean value. The following raises an error:
In [167]: bool(pd.NA)
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
<ipython-input-167-5477a57d5abb> in <module>
----> 1 bool(pd.NA)
/pandas/pandas/_libs/missing.pyx in pandas._libs.missing.NAType.__bool__()
TypeError: boolean value of NA is ambiguous
This also means that pd.NA cannot be used in a context where it is evaluated to a boolean, such as if condition: ... where condition can potentially be pd.NA. In such cases, isna() can be used to check for pd.NA or condition being pd.NA can be avoided, for example by filling missing values beforehand.
A similar situation occurs when using Series or DataFrame objects in if statements, see Using if/truth statements with pandas.
NumPy ufuncs
pandas.NA implements NumPy’s __array_ufunc__ protocol. Most ufuncs work with NA, and generally return NA:
In [168]: np.log(pd.NA)
#Out[168]: <NA>
In [169]: np.add(pd.NA, 1)
#Out[169]: <NA>
Warning Currently, ufuncs involving an ndarray and
NAwill return an object-dtype filled with NA values.In [170]: a = np.array([1, 2, 3]) In [171]: np.greater(a, pd.NA) #Out[171]: array([<NA>, <NA>, <NA>], dtype=object)The return type here may change to return a different array type in the future.
See DataFrame interoperability with NumPy functions for more on ufuncs.
Conversion
If you have a DataFrame or Series using traditional types that have missing data represented using np.nan, there are convenience methods convert_dtypes() in Series and convert_dtypes() in DataFrame that can convert data to use the newer dtypes for integers, strings and booleans listed here. This is especially helpful after reading in data sets when letting the readers such as read_csv() and read_excel() infer default dtypes.
In this example, while the dtypes of all columns are changed, we show the results for the first 10 columns.
In [172]: bb = pd.read_csv('data/baseball.csv', index_col='id')
In [173]: bb[bb.columns[:10]].dtypes
#Out[173]:
player object
year int64
stint int64
team object
lg object
g int64
ab int64
r int64
h int64
X2b int64
dtype: object
In [174]: bbn = bb.convert_dtypes()
In [175]: bbn[bbn.columns[:10]].dtypes
#Out[175]:
player string
year Int64
stint Int64
team string
lg string
g Int64
ab Int64
r Int64
h Int64
X2b Int64
dtype: object
Source : .